Job Arrays
Last updated on 2024-05-07 | Edit this page
Overview
Questions
- How can I run a lot of similar jobs?
- How can I use job arrays in conjunction with a CSV file?
Objectives
- Understand the job array syntax
- Understand how to make use of array indices
- Know how to use
sedandreadto parse
Processing Multiple Cases
Now that you feel pretty comfortable with the pi-cpu
program, your supervisor wants you to investigate how the error value of
the tool changes with the number of iterations. This is controlled with
the -n flag.
He’s noted that the tool uses 123,456,789 iterations (approx. 1.2E8), but he would like to know what the accuracy is like for lower number of iterations. He would like you to see how the error value behaves for 1E2, 1E3, 1E4, …, up to 1E8 iterations.
You know that running 7 tests manually is not a big deal, but you decide that you might want to make this easier to use in case the number of tests increase in the future.
You’ve heard about Slurm job arrays which is good at breaking up these “parameter scans”, so you decide to give them a go.
What does a job array do?
The job array functionality is a lightweight mechanism that Slurm
provides to allow users to submit many similar Slurm scripts with a
single sbatch command.
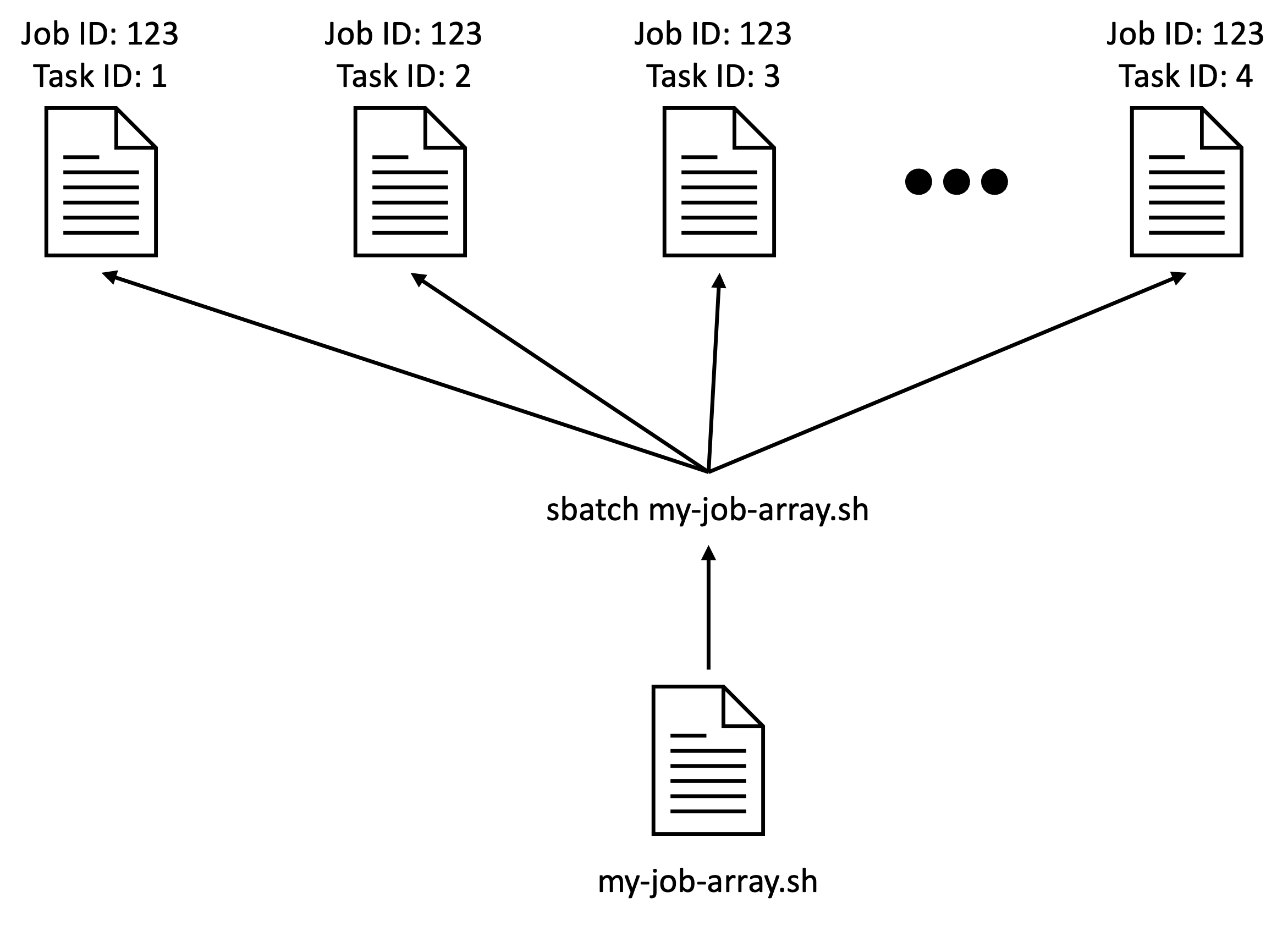
Something similar could be achieved with a for loop
like:
which submits my-script.sh 6 times, and passing the
numbers 1 to 6 to each submission. However, job arrays have some
advantages over this approach:
- job arrays are more self-contained i.e., they do not need additional wrapper scripts or code to submit to Slurm.
- each job array “task” is linked to the same job ID, and can be more
easily queried with
sacctandsqueue.
Why you might prefer the “for loop” approach over job arrays:
- each task in the job array has the same resources - seperate
sbatchcommands allow you to change the resource request. - when the work being done differs significantly between tasks - making it difficult to control the behaviour solely through a “task ID”.
Job Array Syntax
A Slurm script is turned into a job array with the
--array=<range> sbatch option. On Milton,
<range> can be any integer from 0 up to 1000. To
specify a sequential range of values, you can use the
min-max syntax, where min and max
are the minimum and maximum values of the range.
For example, your Slurm script may look like
This script will execute 10 jobs which request the default CPUs and memory. The job will print “hello world” to the job’s output file, and then wait for an hour. When you submit this job, and your job waits in the queue, this array job will look similar to:
OUTPUT
Submitted batch job 11784178OUTPUT
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
11784178_[1-3] regular myjob yang.e PD 0:00 1 (None)where the [1-3] correspond to the indices passed to
--array. Once they start running, the output from
squeue will look similar to:
OUTPUT
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
11784178_1 regular myjob yang.e R 0:00 1 sml-n02
11784178_2 regular myjob yang.e R 0:00 1 sml-n02
11784178_3 regular myjob yang.e R 0:00 1 sml-n02Each job is referred to as a “task” of a single job, each associated with their own index. In the above example, each array task will have a task ID of 1-10.
The --array option can also accept lists of integers and
ranges, seperated by commas. For example, --array=1-3,7 is
acceptable too! This is useful for testing or rerunning only specific
indices.
Each array task will have its own output file. The default naming is
slurm-<jobID>-<taskID>.out. For example, the
above job submission produced:
OUTPUT
slurm-11784178_1.out slurm-11784178_2.out slurm-11784178_3.outCancelling array jobs
When cancelling jobs, you can reference individual array tasks, a range, or a list similar to specifying the jobs to run. For example,
will cancel only array tasks 1, 2, and 3. If you want to cancel all the job tasks at once, you can pass only the job ID.
This will cancel any unfinished or pending array tasks in the queue.
Note that this only works for scancel! No other Slurm
command will accept this format.
Environment Variables in Slurm Jobs
Before you learn more about making use of job arrays, it’s important to know about Slurm job environment variables!
Every time Slurm runs a job for you, Slurm makes a number of environment variables available to you. These environment variables contain information about the job, like the job ID, the job name, or the job’s resources.
A full list of the available environment variables in a job can be
found with man sbatch and scrolling down to
OUTPUT ENVIRONMENT VARIABLES. You can jump to this section
by typing /output environment variables after you’ve opened
the manual page.
Challenge
Try crafting a Slurm script that requests 4 CPUs and 4GB of memory that says hello, and then prints the node it’s running on, the number of CPUs requested, and then the memory requested. An example output would be:
OUTPUT
Hello. I am running on sml-n01
These are the resources I requested:
CPUs: 4
Memory: 4096MBThe Slurm environment variables needed here are
SLURM_CPUS_PER_TASK, SLURM_JOB_NODELIST, and
SLURM_MEM_PER_NODE.
To produce the output in the challenge, your script needs to look similar to:
BASH
#!/bin/bash
#SBATCH --cpus-per-task=4
#SBATCH --mem=4G
echo "Hello. I am running on ${SLURM_JOB_NODELIST}"
echo "These are the resources I requested:"
echo "CPUs: ${SLURM_CPUS_PER_TASK}"
echo "Memory: ${SLURM_MEM_PER_NODE}MB"These parameters can be useful when you would like to automatically
pass the Slurm resource request to a command in your script. For
example, I can make pi-cpu use however many CPUs I’ve
requested automatically by using the `SLURM_CPUS_PER_TASK environment
variable inside a job script:
But be careful: some Slurm environment variables only have a value if
you’ve set it as a flag. For example, in the above example script,
SLURM_CPUS_PER_TASK has a value because we supplied
--cpus-per-task. But if you didn’t set
--cpus-per-task, the corresponding environment variable
would be empty!
The SLURM_ARRAY_TASK_ID Environment Variable
Now back to Slurm job arrays!
While skimming through the manual pages, you might’ve noticed that
one of the environment variables listed is
SLURM_ARRAY_TASK_ID. This variable will store the ID of
each task in a job array, which can be used to control what each task
does.
As practice, we can take the Slurm script we wrote at the beginning and modify it a little:
BASH
#!/bin/bash
#SBATCH --job-name=myjob
#SBATCH --array=1-3
echo "hello world from task ${SLURM_ARRAY_TASK_ID}"This script should now print
hello world from task <N> where N is
1-3.
OUTPUT
Submitted batch job 11784442And once all the job tasks have completed, we should be able to check the output of each task:
OUTPUT
hello world from task 1
hello world from task 2
hello world from task 3Which is great! Now we just have to figure out how to make use of
these task IDs. Remember: our goal is to execute
pi-cpu -n <iter> where iter is 1E2, 1E3,
…, 1E8.
One might think of using the IDs directly, for example, instead of
#SBATCH --array=1-3, maybe we could try something like
But if we add this to our example array script and try to submit it, we get the error:
ERROR
sbatch: error: Batch job submission failed: Invalid job array specificationAnd unfortunately this is because Slurm on Milton has been configured to accept a max job array index of 1000.
So, we have to figure out another way! A common alternative is to use
a file to store the values we want the job array to scan over. In our
case, we can put together a file with a line for each number of
iterations we want the job array to scan over. Let’s call it
iterations.txt, which contains:
100
1000
10000
100000
1000000
10000000
100000000But how do we use the job array index to retrieve each of these
lines? We can use the readarray command line utility.
readarray is a neat tool that can split text into a Bash
array. Executing the command will start an interactive prompt. To exit,
press Ctrl+D.
Your text gets saved into the MAPFILE array variable
(each index corresponds to a line):
OUTPUT
line1: this, line2: is an, line3: exampleA couple things to note about referencing elements in bash arrays: *
Array indices start at 0 * ${} are on the outside of
array[index].
Instead of supplying it input manually, we can pass it a file using redirection:
OUTPUT
100 1000 10000 100000 1000000 10000000 100000000The @ symbol means “all the elements in the array”.
Instead of saving the array into MAPFILE, we can pass a
variable name to readarray and it will save the array into
that variable instead e.g.:
OUTPUT
100 1000 10000 100000 1000000 10000000 100000000Can you start to see how we might combine the Slurm array index and
our bash array to pass different iteration values to
pi-cpu?
Let’s first test that we know how to properly combine the
SLURM_ARRAY_TASK_ID environment variable together with
readarray. Take your script and modify it:
BASH
#!/bin/bash
#SBATCH --job-name=myjob
#SBATCH --array=0-2
echo "hello world from task ${SLURM_ARRAY_TASK_ID}"
readarray niterations < iterations.txt
echo "I will run ${niterations[$SLURM_ARRAY_TASK_ID]}!"Note that we’ve changed the array task range from 1-3 to 0-2 since the bash array is 0-indexed.
Let’s submit this script to confirm that we’ve used `readarray`` correctly.
OUTPUT
Submitted batch job 11784737Because we’ve only passed the range 0-1 to the
--array option, we should expect to only see outputs for
the first 3 rows in iterations.txt:
OUTPUT
hello world from task 0
I will run 100
!
hello world from task 1
I will run 1000
!
hello world from task 2
I will run 10000
!Which demonstrates that we’ve taken the first 3 lines of
iterations.txt correctly! But there’s something wrong… the
exclamation marks on the next line instead of at the end of the number!
This is because readarray keeps newline characters when
parsing the file. To turn off this behavior we need to add the
-t option, so our readarray command
becomes:
We can now make use of these values by passing the number of
iterations to the pi-cpu command. We also need to change
the array range to pull all the lines of the iterations.txt
file. Change the array range and add the pi-cpu command to
your script:
BASH
#!/bin/bash
#SBATCH --job-name=myjob
#SBATCH --array=0-6
#SBATCH --cpus-per-task=4
echo "hello world from task ${SLURM_ARRAY_TASK_ID}"
readarray -t niterations < iterations.txt
echo "I will run ${niterations[$SLURM_ARRAY_TASK_ID]}!"
srun ./pi-cpu -p ${SLURM_CPUS_PER_TASK} -n ${niterations[$SLURM_ARRAY_TASK_ID]}You will also need to ensure that --cpus-per-task is
provided here, as without that option, SLURM_CPUS_PER_TASK
doesn’t get set either.
We can then submit the script:
OUTPUT
$ cat slurm-11913780_*.out
hello world from task 0
I will run 100!
Result: 3.1600000000000 Error: 0.0184072589874 Time: 0.0003s
... skipped output
hello world from task 1
I will run 1000!
Result: 3.0480000000000 Error: -0.0935927410126 Time: 0.0004s
... skipped output
hello world from task 2
I will run 10000!
Result: 3.1344000000000 Error: -0.0071927410126 Time: 0.0056s
... skipped output
hello world from task 3
I will run 100000!
Result: 3.1431600000000 Error: 0.0015672589874 Time: 0.0013s
... skipped output
hello world from task 4
I will run 1000000!
Result: 3.1421880000000 Error: 0.0005952589874 Time: 0.0100s
... skipped output
hello world from task 5
I will run 10000000!
Result: 3.1419520000000 Error: 0.0003592589874 Time: 0.0936s
... skipped output
hello world from task 6
I will run 100000000!
Result: 3.1414855200000 Error: -0.0001072210126 Time: 0.9331s
... skipped outputAnd now you can see the magnitude of the error, and the speed decreasing as we increase the number of iterations.
Multi-column data
So far you’ve only needed one column of data (the number of iterations) to investigate accuracy with number of iterations. But, in many cases you might be interested in varying multiple variables with each array task.
We can do this by adding another column to our data. Create a new file called iter-cpu.txt with the content:
iters cpus
100 2
100 4
1000 2
1000 4
10000 2
10000 4The first row is now a header and rows 2-7 contains the data we’ll
use in our job arrays. Here, we’re going to vary the value passed to
-p.
We can still execute readarray on this file:
OUTPUT
iters cpusBut the columns don’t get split!
So how do we split the columns? cut is a tool you might
be aware of (for example from an introductory
course). But, this time, lets split the lines of text using the
read utility.
We can use the “heredoc” operator <<< to pass
text to the read command:
OUTPUT
hello worldBy default, read saves what we send it to the variable
REPLY. Note that unlike readarray,
REPLY is not an array. We can choose the variable to save
our input to by giving read a variable name:
OUTPUT
hello worldread is also useful because if we pass it more variable
names, it will split the words into each of thoses variables. So, if we
add another variable name to our previous read command:
OUTPUT
myvar1: hello, myvar2: worldparsing theiter-cpu.txtfile
We now know how to split lines of a file into an array using
readarray, as well as splitting strings with spaces into
individual words!
See if you can apply this to our current case: try and get
the first line from iter-cpu.txt and save
the first column into niterations and the second column
into ncpus bash variables
As shown already we can save iter-cpu.txt into an array
using
This gives us an array, rowdata, where each element is a
line of text from iter-cpu.txt. To split the first line of
the file into variables niterations and
ncpus:
${rowdata[0]} is referencing the first element of
rowdata, and therefore the first line of text in
iter-cpu.txt. read will take this the row
data, and split the two words into niterations and
ncpus variables.
Yes we do! While we have the same number of rows in
iter-cpu.txt, the first row is headers. If we passed those
headers to pi-cpu instead of integer values, the program
would fail. We need to ensure only array tasks 1-6 are being run
(remember: the 0th index is the first row in the file i.e., the
headers).
Be aware of this when you write your own files and job array scripts!
We first need to modify the readarray line to read the
correct file, and to use an appropriately named array variable:
We then need to split rowdata further into
niterations and ncpus. To do this, we will
adapt what we wrote for “Parsing the iter-cpu.txt file”
challenge. Instead of reading the first “0th” element of
rowdata, we’ll use the SLURM_ARRAY_TASK_ID
environment variable:
And finally, we can add niterations and
ncpus to our echo and pi-cpu execution
command!
BASH
echo "I will run $niterations iterations and with $ncpus CPUs!"
srun pi-cpu -p $ncpus -n $niterationsOur final script should look something like:
BASH
#!/bin/bash
#SBATCH --job-name=myjob
#SBATCH --array=1-6
#SBATCH --cpus-per-task=4
echo "hello world from task ${SLURM_ARRAY_TASK_ID}"
readarray -t rowdata < iter-cpu.txt
read niterations ncpus <<< ${rowdata[$SLURM_ARRAY_TASK_ID]}
echo "I will run $niterations with $ncpus CPUs!"
srun pi-cpu -p $ncpus -n $niterationsControlling output
When using --output and --error flags with
sbatch, you can make use of the %A variable
which refers to the parent job ID, and %a, which refers to
the job task. Using %j will use a different job ID for each
task.
Exception handling
Sometimes, you might find that you want to execute your job script
from the command line, instead of submitting it to Slurm. In this case,
your job likely won’t have a SLURM_ARRAY_TASK_ID
environment variable set. In this case, you will want to make sure the
necessary check is present and for the script to exit appropriately OR
set a reasonable default.
BASH
if [ -z $SLURM_ARRAY_TASK_ID ]
then
echo "This script needs to be submitted as a Slurm script!"
exit 1
fi[ -z $SLURM_ARRAY_TASK_ID ] returns 0 if
SLURM_ARRAY_TASK_ID is NOT set. You can replace the
echo and exit statement with a reasonable
default instead e.g., export SLURM_ARRAY_TASK_ID=1.
Different delimiters (optional)
Here we showed how to split text that is seperated by spaces. e.g.
But different delimiters can be used using the following syntax:
IFS is a special environment variable used by bash to
determine how to split strings. By setting it to a different
character(s), you can control how read splits your string.
What should you set IFS to to get the following output from
echo "word1: $word1, word2: $word2"?
word1: hello , word2: orldword1: , word2: ello world
Key Points
- Slurm job arrays are a great way to parallelise similar jobs!
- The
SLURM_ARRAY_TASK_IDenvironment variable is used to control individual array tasks’ work - A file with all the parameters can be used to control array task parameters
-
readarrayandreadare useful tools to help you parse files. But it can also be done many other ways!
